Difference between revisions of "Main Page"
From SEED
| Line 27: | Line 27: | ||
{| | {| | ||
| + | |- | ||
|Remember to cite SEED: Větrovský, T. and P. Baldrian (2013). "Analysis of soil fungal communities by amplicon pyrosequencing: current approaches to data analysis and the introduction of the pipeline SEED." Biology and Fertility of Soils 49: 1027-1037. | |Remember to cite SEED: Větrovský, T. and P. Baldrian (2013). "Analysis of soil fungal communities by amplicon pyrosequencing: current approaches to data analysis and the introduction of the pipeline SEED." Biology and Fertility of Soils 49: 1027-1037. | ||
|} | |} | ||
Revision as of 12:57, 12 March 2015
|
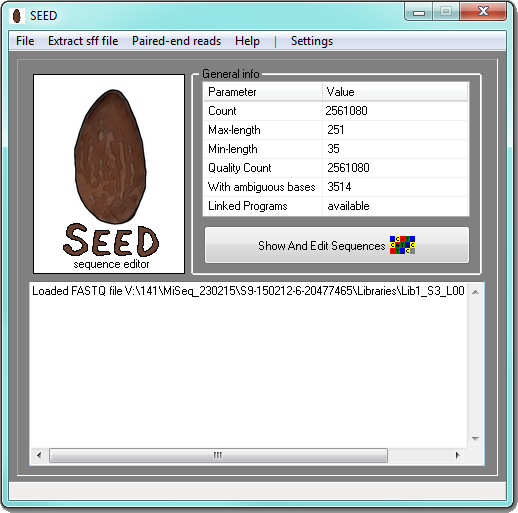
SEED is a simple sequence editor and pipeline for processing of nucleotide sequences in the FASTA and FASTQ format. SEED allows simple grouping of sequences based on the search for sequence motifs (e.g. primers, pyrosequencing tags) and a batch processing of the resulting groups. It also supports the connections to external tools for sequence processing and analysis like Mafft, Mothur, Uclust, Usearch, ITSx or Blast and can be used for further processing, visualization, management and analysis of their results. The SEED is especially designed to provide the environment for a sequential analysis of the sequences of PCR amplicons obtained by 454-pyrosequencing or Illumina.
| Remember to cite SEED: Větrovský, T. and P. Baldrian (2013). "Analysis of soil fungal communities by amplicon pyrosequencing: current approaches to data analysis and the introduction of the pipeline SEED." Biology and Fertility of Soils 49: 1027-1037. |
- See the workflows - See some analysis examples.
- See the list of functions - Feel free to take a look around at what SEED can do for you.
- SEED main page
Current version of SEED is 1.2.3