The study of structure and function of 14-3-3 proteins and their complexes
Our research team has been studying the 14-3-3 proteins which are highly conserved regulatory molecules found in all eukaryotes. 14-3-3 proteins have the ability of binding the functionally different signal proteins, including kinases, fosfatases and transmembrane receptors by changing their function. Through the functional modulation of a wide range of binding partners, 14-3-3 proteins are involved in many processes, including cell cycle regulation, metabolism control, apoptosis, and control of gene transcription. More than 300 proteins have been described as binding partners till now. We employ both biophysical (fluorescence spectroscopy, analytical ultracentrifugation, SAXS, mass spectrometry, isothermal titration calorimetry, X-ray crystallography, protein structure modeling, etc.) and biochemical (recombinant protein expression, site-directed mutagenesis, enzyme kinetics) approaches to understand the details of how the activity and function of protein-protein complexes are regulated.
Research Projects
The 14-3-3 proteins are a family of regulatory molecules, which specifically bind to phosphoserine (or phosphothreonine)- containing motifs (pSer/pThr) in a sequence-specific manner. Through these binding reactions, the 14-3-3 proteins play key regulatory roles in signal transduction, cell cycle control, metabolism control and apoptosis. More than 200 14-3-3 binding partners have been reported so far and some of them play prominent roles in cancer development (e.g. transcription factors p53 and FOXO), neurodegeneration (e.g. Tau protein, ASK1 kinase), cardiovascular diseases (e.g. RGS proteins, phosducin) or inflammation (e.g. NFkB, ASK1 kinase). However, the detailed mechanisms of the14-3-3 protein-mediated regulations are mostly elusive, mainly due to the lack of structural data.
Main goal of our research is a mechanistic understanding of the 14-3-3 protein function in the regulation of selected 14-3-3 protein binding partners. In recent years we have been studying the 14-3-3 protein-mediated regulation of forkhead transcription factor FOXO4, tyrosine hydroxylase, regulator of G-protein signaling RGS3 or enzyme neutral trehalase. Our current projects are focused on regulation of ubiquitin ligase Nedd4-2, FOXO-DBD, caspase-2 and proten kinases: CaMKK2, ASK1 and DAPK2.
Nedd4-2
Ubiquitin E3 protein ligase Nedd4-2 belongs to the physiological regulators of the epithelial sodium channel, which is responsible for sodium reabsorption by the epithelia lining the distal part of the kidney tubule. The mutations in the Nedd4-2 gene are associated with the developmental disorders, hypertension and epilepsy. The function of Nedd4-2 is negatively regulated by the presence of vasopressin or aldosterone which induces the phosphorylation of the specific serine residues through signaling cascades of Sgk1 and PKA. This phosphorylation enables the binding of 14-3-3 proteins, thus preventing the binding between Nedd4-2 and its substrate. Structural basis of this 14-3-3-dependent regulation of Nedd4-2 is still unresolved. To elucidate this regulatory mechanism we propose to: i) investigate the 14-3-3 binding specificity for Nedd4-2; ii) perform the biophysical and structural characterization of the 14-3-3:Nedd4-2 complex; and iii) map its binding interface. The proposed research is expected to provide the structural insight into the 14-3-3-dependent regulation of Nedd4-2. This project was funded by Czech Science Foundation (Projects 23-04686S and 20-00058S).
Results were published in: Janosev et al. (2025) Nature Communications, Joshi et al. (2022) Biophysical Journal, Pohl et al. (2021) Communications Biology.
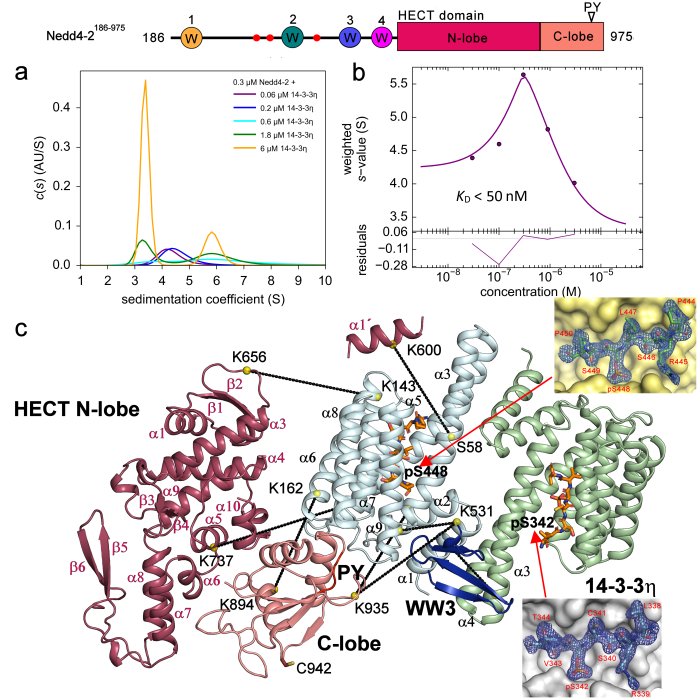
Structural analysis of the complex between Nedd4-2 and 14-3-3 protein by integrative structural biology approach. a)+b) Sedimentation velocity analytical ultracentrifugation (SV-AUC) measurements with the estimation of the apparent dissociation constant of the complex. c) SAXS-based structural modeling with the crystal structures of the 14-3-3 binding sites of Nedd4-2 (Pohl et. al (2021) Communications Biology).
FOXO and p53
The overall goal of this project is to understand the molecular basis of the interaction between Forkhead box O (FOXO) transcription factors and tumor suppressor p53. These proteins not only regulate diverse signaling pathways to control cell cycle, apoptosis and metabolism, but also play a crucial role in aging and their misregulation has been shown to be linked to several pathologies including cancer. Recent studies have shown that p53 and FOXO transcription factors physically interact and that this interaction plays an essential role in drug resistance and tissue aging. However, many details concerning the interaction between p53 and FOXO are still unresolved. To better understand this protein-protein interaction, we propose to perform biophysical and structural analysis of complexes between p53 and FOXO proteins using analytical ultracentrifugation, NMR, and H/D exchange and chemical cross-linking coupled to mass spectrometry. The proposed research will enable to better understand the molecular basis of the interaction between p53 and FOXO proteins. This project was funded by Czech Science Foundation (Project 21-02080S).
Results were published in: Kohoutova et al. (2025) Nature Communications, Kohoutova et al. (2022) ACS Omega, Mandal et al. (2022) Protein Science.
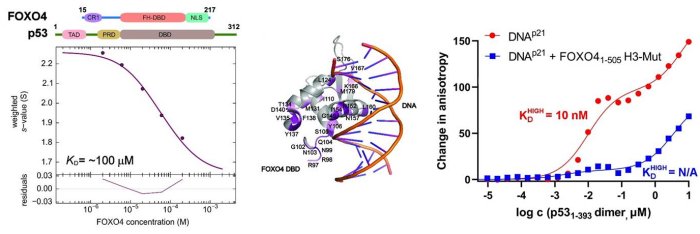
Left, sedimentation velocity analytical ultracentrifugation analysis of interaction between FOXO4 and p53. Middle, chemical shift perturbations obtained from 1H-15N HSQC spectra of 15N-labeled FOXO4 in the presence of p53 mapped onto the crystal structure of the FOXO4 DBD:DNA complex. Right, fluorescence anisotropy measurements showing that the complex formation reduces the DNA-binding affinity of p53 (Mandal et al. (2022) Protein Science).
Protein kinases ASK1, CaMKK2 and DAPK2
14-3-3 proteins, a family of highly conserved dimeric proteins ubiquitously expressed in all eukaryotic cells, regulate the function of several hundred partner proteins including many disease-relevant protein kinases. However, our understanding of how the 14-3-3 proteins regulate the activity of these protein kinases is, in most cases, very limited mainly due to the lack of structural data. This proposal aims to investigate the molecular mechanisms underlying the 14-3-3 protein-mediated regulation of physiologically important serine/threonine protein kinases: Apoptosis signal-regulating kinase 1 (ASK1), Ca2+/calmodulin-dependent protein kinase kinase (CaMKK) and Death-associated protein kinase 2 (DAPK2). 14-3-3 proteins have been shown to inhibit the activity of these kinases but mechanisms governing these processes are unclear. Therefore, we propose to perform structural and functional analysis of 14-3-3 protein complexes with protein kinases ASK1, CaMKK and DAPK2 using an integrative approach based on protein crystallography, SAXS, mass spectrometry and fluorescence spectroscopy. This project was funded by Czech Science Foundation (Project 19-00121S).
Results were published in: Horvath et al. (2021) Communications Biology; Obsilova et al. (2020) International Journal of Molecular Sciences; Lentini Santo et al. (2020) ACS Chemical Biology; Psenakova et al. (2020) FEBS Journal
Caspase-2
Caspase-2 (C2), a cysteine-dependent and aspartate-specific intracellular protease, has multiple roles in the DNA damage response, cell cycle regulation and tumor suppression. C2 functions as a central coordinator between the cell metabolism and apoptosis and its function is regulated by phosphorylation at several Ser residues. Phosphorylated procaspase-2 (proC2) binds to the 14-3-3 protein and this interaction blocks proC2 activation through an unknown mechanism. To elucidate this regulatory mechanism we propose to: i) identify sites responsible for the 14-3-3 protein binding to proC2, ii) perform biophysical characterization of the 14-3-3:proC2 complex using analytical ultracentrifugation, iii) map the binding interface of the 14-3-3:proC2 complex and perform its structural analysis using hydrogen-deuterium exchange coupled to MS, SAXS and protein crystallography. The proposed research is expected to provide the structural insight into the 14-3-3-dependent regulation of C2. This project was funded by Czech Science Foundation (Project 17-00726S).
Results were published in: Kalabova, Filandr et al. (2020) FEBS Journal; Smidova, Alblova, Kalabova et al. (2018) FEBS Journal; Kalabova et al. (2017) BBRC.
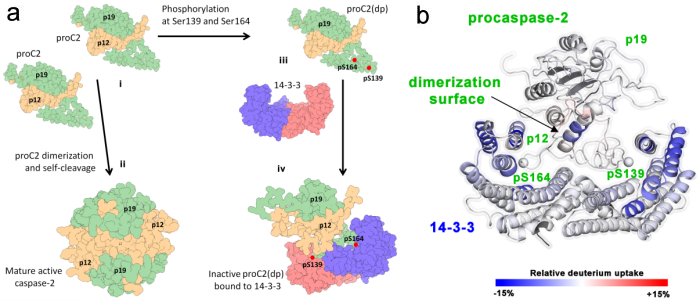
a) Schematic model of regulation of caspase-2 by 14-3-3 protein, b) HDX-MS reveals the biniding interface of procaspase-2 and 14-3-3 (Kalabova, Filandr et. al (2020) FEBS Journal).
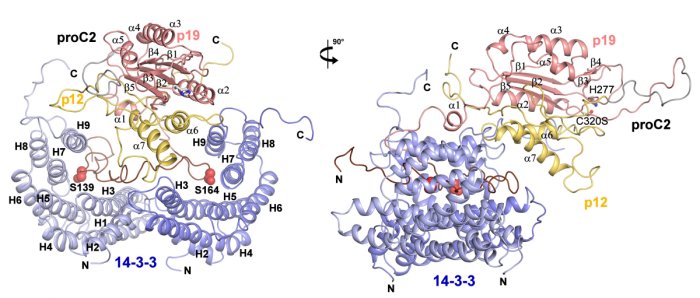
SAXS-based structural modeling of the complex between procaspase-2 and 14-3-3 protein (Smidova, Alblova, Kalabova et. al (2018) FEBS Journal).
FOXO3-DBD
FOXO transcription factors control apoptosis, stress resistance and longevity in mammalian cells. Although the members of the FOXO family act as tumor suppressors in some cell types, emerging evidence suggests that FOXO3 also contributes to tumor stem cell renewal, immune suppression, metastases and chemotherapy resistance in certain cancer types. By a combined in silico / cell biological screening approach several small, drug-like compounds that interact with the DNA-binding domain of FOXO3 and efficiently inhibit its transcriptional activity have been identified. The main aim of this project is to define the structural basis for the interaction between compounds and FOXO3 by NMR spectroscopy, design and develop derivatives with improved properties regarding solubility and affinity, analyze compound-dependent inhibition of target recognition by FOXO3 in vitro and in vivo, study the effects of compounds on FOXO3-induced cancer cell survival in 3D cell culture and in vivo. This project is a collaboration with the group of prof. Michael J. Ausserlechner from Medical University Innsbruck, Innsbruck, Austria, and it was funded by Czech Science Foundation (Project 17-33854L).
Results were published in: Hagenbuchner, Obsilova et.al. (2019) eLife; Psenakova, Kohoutova et. al. (2019) Cells
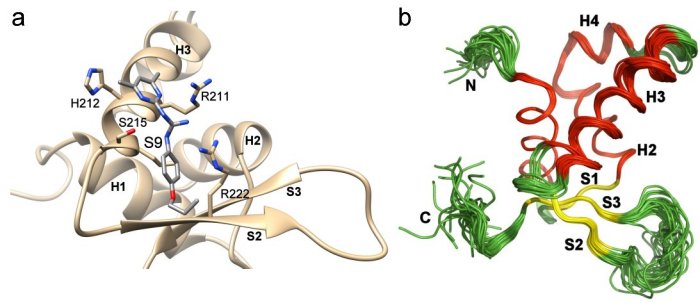
a) Compounds S9 blocks the DNA binding surface of Forkhead transcription factor FOXO3. The figure shows the structural model of the DNA-binding domain of FOXO3 with bound compound S9 based on data from NMR measurements and docking simulations (Hagenbuchner, Obsilova et al. (2019) eLife), b) Solution structure of the free FOXO1 DNA binding domain (Psenakova, Kohoutova et al. (2019) Cells).
Molecular basis of the 14-3-3 protein-dependent activation of yeast neutral trehalase Nth1
We deciphered the mechanism by which the 14-3-3 protein regulates the function of yeast neutral trehalase Nth1. Neutral trehalase Nth1 catalyzes the hydrolysis of disaccharide trehalose and represents one of the key enzymes in the energetic metabolism of yeast. We solved the crystal structures of several forms of neutral trehalase Nth1 and deciphered the changes which occur upon the ligand and 14-3-3 protein binding. We showed that the catalytic activity of Nth1 is enabled by the proper 3D configuration of Nth1´s catalytic and calcium-binding domains relative to each other, which stabilizes the flexible part of the active site required for catalysis. This finalized our long-lasting research focused on understanding of the mechanism of yeast neutral trehalase regulation. Solved atomic structure of the 14-3-3 protein complex with neutral trehalase Nth1 shows the ability of the 14-3-3 protein to modulate the structure of a multidomain enzyme and to function as an allosteric regulator. It is the second solved structure of 14-3-3 protein complex with the fully active enzyme. Comparison of the 14-3-3:Nth1 complex structure with those of other 14-3-3 complexes highlights the ability of 14-3-3 to modulate the structure and function of many client proteins important for the regulation of key cell processes. This project was funded by Czech Science Foundation (Project P207/11/0455).
Results were published in: Alblova et al. (2017) PNAS; Kopecka et al. (2014) JBC; Macakova et al. (2013) BBA-Gen. Subjects; Veisova et al. (2012) Biochemical Journal.
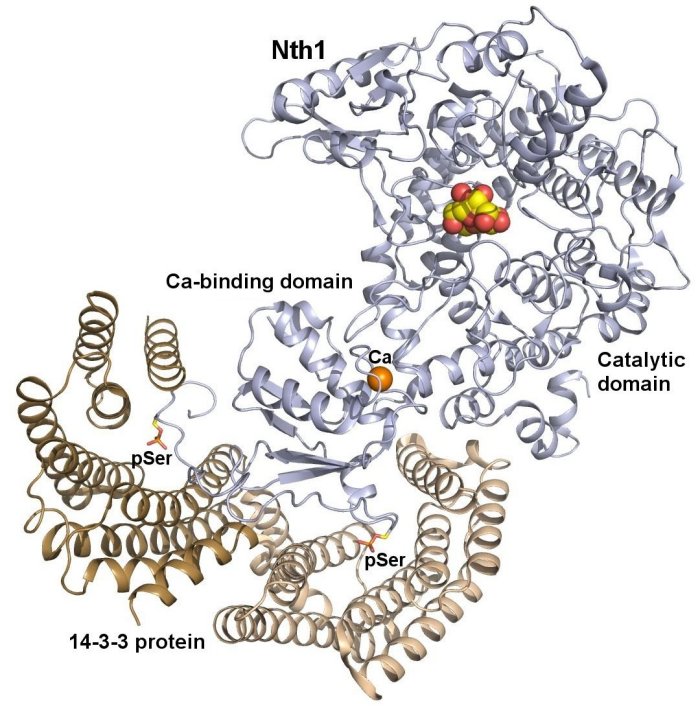
Crystal structure of the complex between phosphorylated Nth and Bmh1 (yeast 14-3-3 protein) (PDB 5N6N). The protomers of the Bmh1 homodimer are shown in yellow and brown. Nth1 is shown in blue. The phosphorylated Ser60 and Ser83 are shown as sticks. The calcium ion is shown in orange. Structural analysis revealed that the binding of phosphorylated Nth1 by 14-3-3 triggers Nth1’s activity by enabling the proper 3D configuration of Nth1’s catalytic and calcium-binding domains relative to each other, thus stabilizing the flexible part of the active site required for catalysis (Alblova et al. (2017) PNAS USA).

